|
§Note: The mechanisms are very different in prokaryotic and eukaryotic organisms - they can also vary between different species and even for different genes! The 3' UTR is simpler to identify - it is typically† everything after the first in frame stop codon and before the polyadenylation signal (where the polyA tail gets added. I also encourage you to look at some of the references for that section, which will help give you more detail on this high complex process that is still being actively studied. This is covered in a bit more detail in another article: (In fact, codons other than AUG are sometimes used as start codons!) These sequences are bound by proteins that help guide the ribosome to assemble at the correct place to start translation. The short answer to that is that the sequence of the mRNA around a potential start codon influences whether or not it will be used§. The interesting question is how does the ribosome know which start codon to start with? Here you will find options to view and activate subscriptions, manage institutional settings and access options, access usage statistics, and more.The 5' UTR is everything 5' of the start codon. If you believe you should have access to that content, please contact your librarian.įor librarians and administrators, your personal account also provides access to institutional account management. The institutional subscription may not cover the content that you are trying to access. Oxford Academic is home to a wide variety of products. View the institutional accounts that are providing access.View your signed in personal account and access account management features.Some societies use Oxford Academic personal accounts to provide access to their members.Ĭlick the account icon in the top right to: See below.Ī personal account can be used to get email alerts, save searches, purchase content, and activate subscriptions. Some societies use Oxford Academic personal accounts to provide access to their members. If you do not have a society account or have forgotten your username or password, please contact your society. Do not use an Oxford Academic personal account. When on the society site, please use the credentials provided by that society.If you see ‘Sign in through society site’ in the sign in pane within a journal: Many societies offer single sign-on between the society website and Oxford Academic. 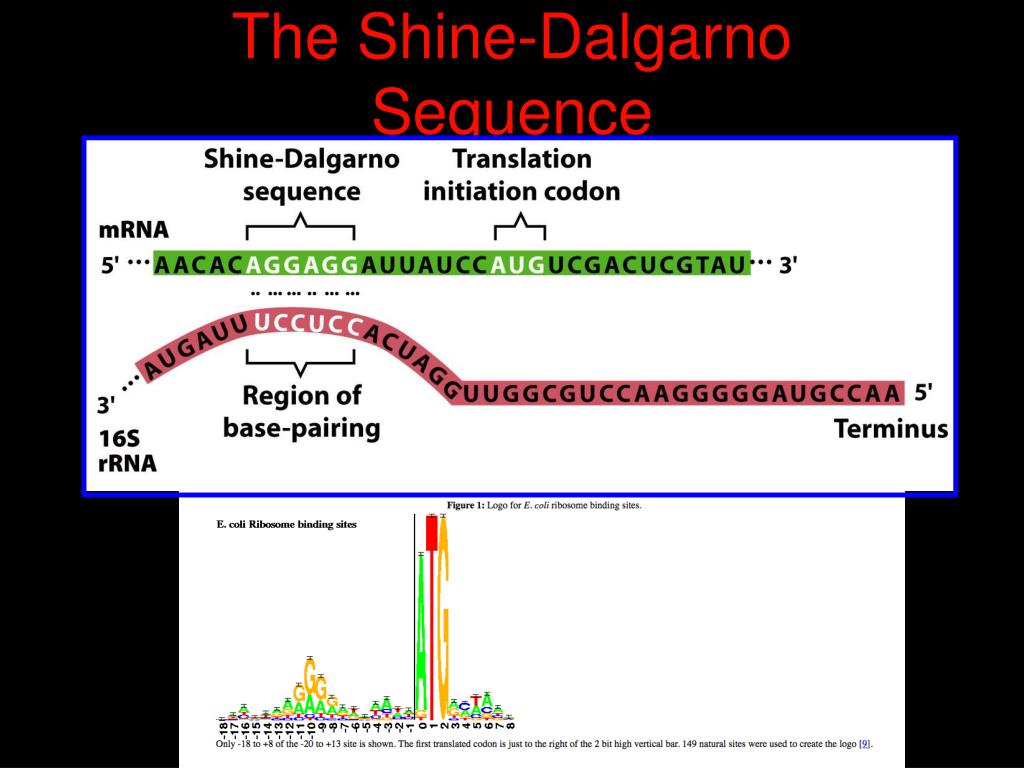
Society member access to a journal is achieved in one of the following ways: If you cannot sign in, please contact your librarian. If your institution is not listed or you cannot sign in to your institution’s website, please contact your librarian or administrator.Įnter your library card number to sign in. Following successful sign in, you will be returned to Oxford Academic.When on the institution site, please use the credentials provided by your institution.Select your institution from the list provided, which will take you to your institution's website to sign in.Click Sign in through your institution.Shibboleth / Open Athens technology is used to provide single sign-on between your institution’s website and Oxford Academic. 
This authentication occurs automatically, and it is not possible to sign out of an IP authenticated account.Ĭhoose this option to get remote access when outside your institution. Typically, access is provided across an institutional network to a range of IP addresses. If you are a member of an institution with an active account, you may be able to access content in one of the following ways: Get help with access Institutional accessĪccess to content on Oxford Academic is often provided through institutional subscriptions and purchases. Since aligned spacing corresponds naturally to the spacing between the 3′-end of the 16s rRNA and the P-site, we conclude that there is a unique optimal aligned SD– AUG spacing in the absence of Measurements of protein expression demonstrated an optimal aligned spacing of 5 nt for both series. We thus undertook a systematic study by inserting two series of synthetic RBSs of varying spacing and SD sequence into a plasmid vector containing the chloramphenicol acetyltransferase gene.Ĭare was taken not to introduce any secondary structure. Complications involving the definition of the spacing as well as secondary structures have obscured matters. It is not as clear, however, whether there is a unique optimal spacing. It is now clear that the SD sequence is important for identification of the translation initiation site on the mRNA by the ribosome, and that as a result, the spacing between the SD and the initiation codon strongly affects translational efficiency (1). The prokaryotic mRNA ribosome binding site (RBS) usually contains part or all of a polypurine domain UAAGGAGGU known as the Shine – Dalgarno (SD) sequence found just 5′ to the translation initiation codon.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
